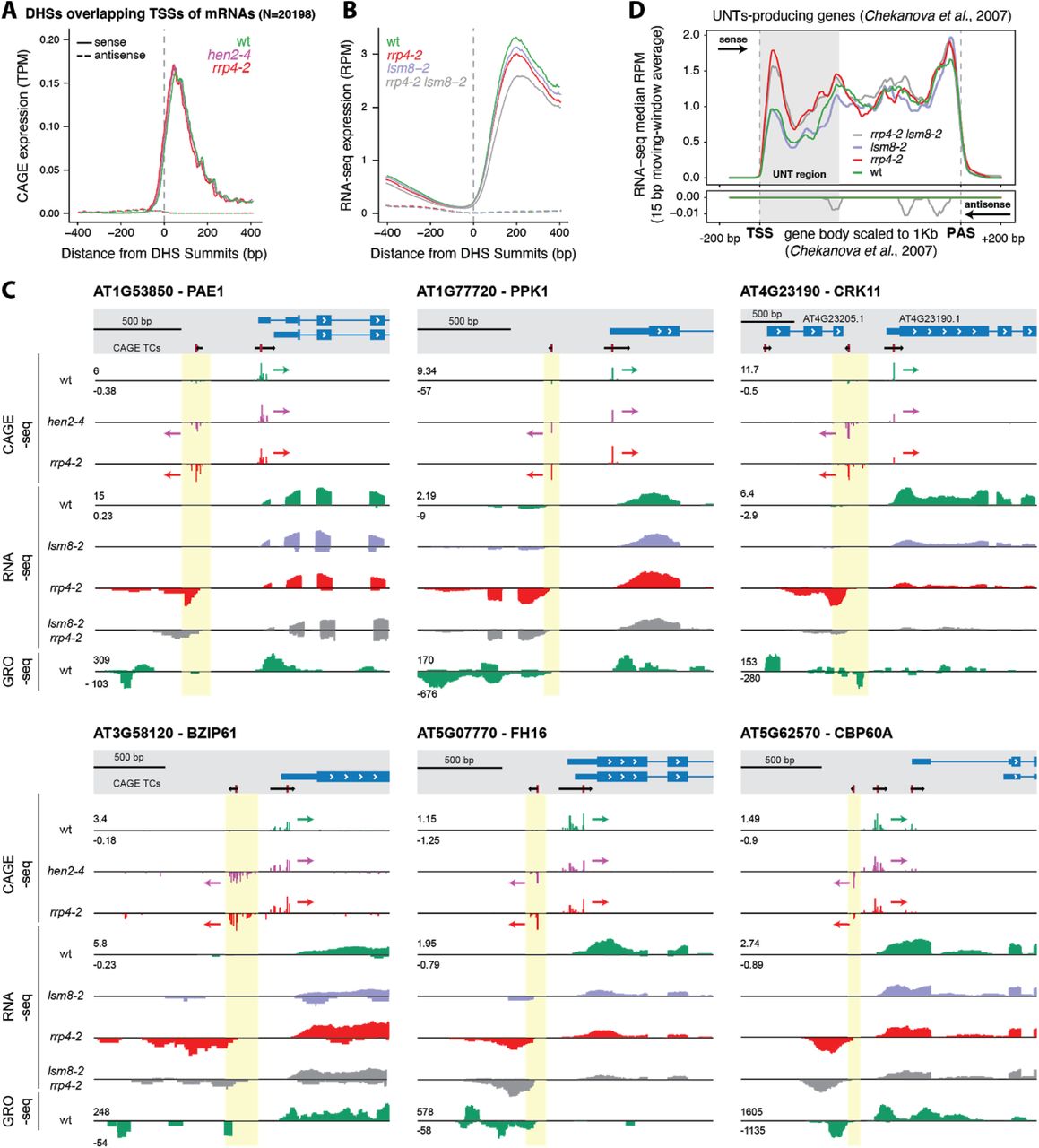
Characterization of Arabidopsis thaliana promoter bidirectionality and antisense RNAs by depletion of nuclear RNA decay enzymes | bioRxiv
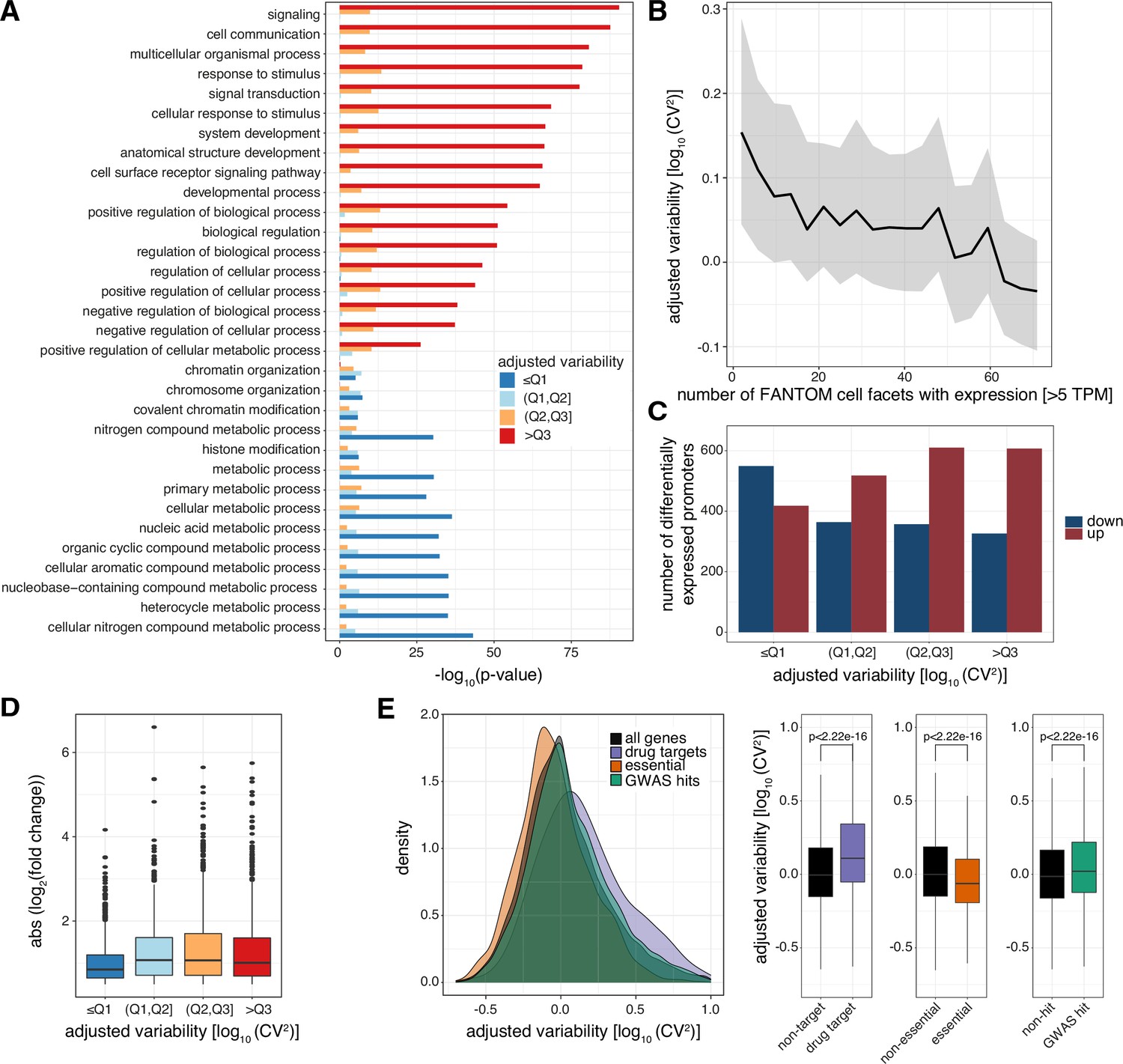
Promoter sequence and architecture determine expression variability and confer robustness to genetic variants | eLife

Cap analysis of gene expression reveals alternative promoter usage in a rat model of hypertension | Life Science Alliance
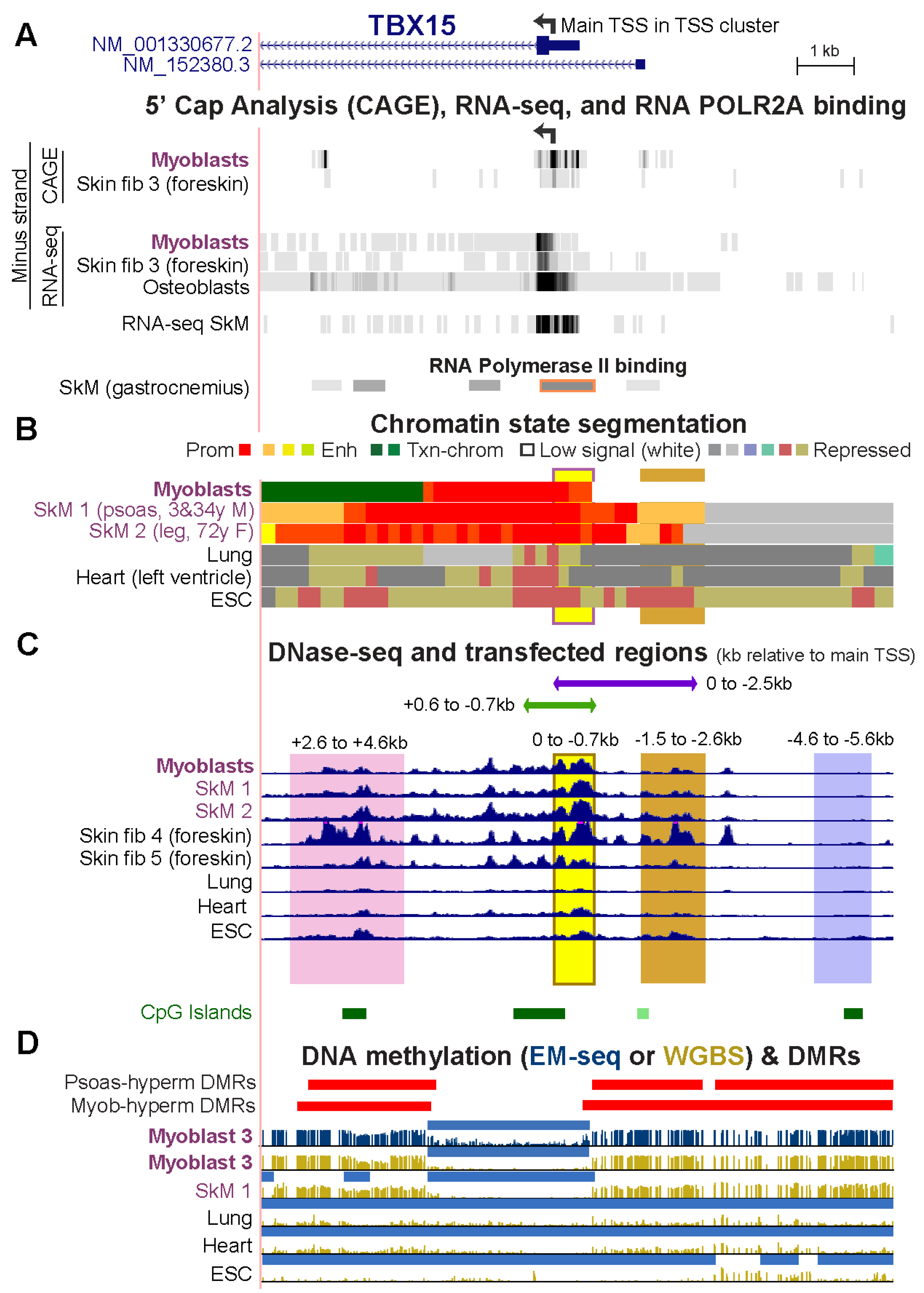
Epigenomes | Free Full-Text | Promoter-Adjacent DNA Hypermethylation Can Downmodulate Gene Expression: TBX15 in the Muscle Lineage

CAGE-seq characterization of alternative promoter usage and full-length... | Download Scientific Diagram
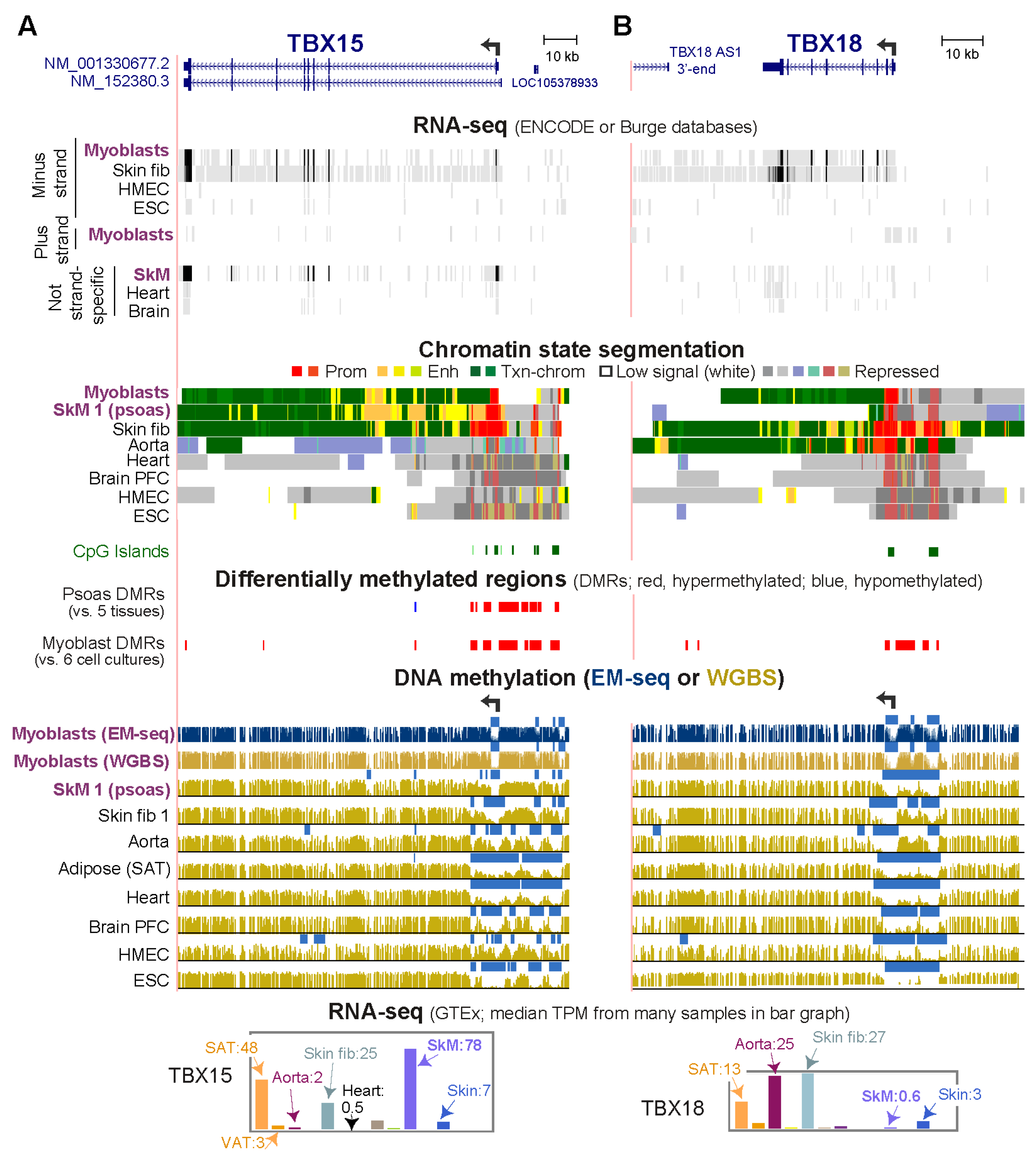
Epigenomes | Free Full-Text | Promoter-Adjacent DNA Hypermethylation Can Downmodulate Gene Expression: TBX15 in the Muscle Lineage
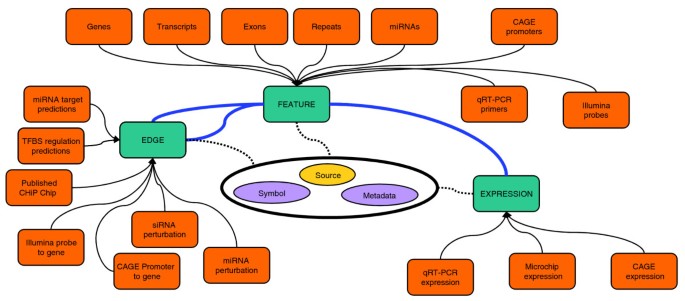
FANTOM4 EdgeExpressDB: an integrated database of promoters, genes, microRNAs, expression dynamics and regulatory interactions | Genome Biology | Full Text

Alternative promoters in CpG depleted regions pervasively account for epigenetic misregulation of cancer transcriptomes | bioRxiv